the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Is Colobus guereza gallarum a valid endemic Ethiopian taxon?
Dereje Tesfaye
Nils C. Stenseth
Afework Bekele
Aemro Mekonnen
Steve Doeschner
Anagaw Atickem
Christian Roos
Black-and-white colobus (Colobus guereza Rüppell, 1835) are arboreal Old World monkeys inhabiting large parts of the deciduous and evergreen forests of sub-Saharan Africa. Two of the eight subspecies of Colobus guereza are endemic to Ethiopia: C. g. gallarum and C. g. guereza. However, the validity of the Ethiopian taxa is debated and observed morphological differences were attributed to clinal variation within C. g. guereza. To date, no molecular phylogeny of the Ethiopian guerezas is available to facilitate their taxonomic classification. We used mitochondrial DNA markers from 94 samples collected across Ethiopia to reconstruct a phylogeny of respective mitochondrial lineages. In our phylogenetic reconstruction, augmented by orthologous sequence information of non-Ethiopian black-and-white colobus from GenBank, we found two major Ethiopian mitochondrial clades, with one being largely congruent with the distribution of C. g. guereza. The second clade was found only at two locations in the eastern part of the putative range of C. g. gallarum. This second lineage clustered with the lowland form, C. g. occidentalis, from central Africa, whereas the C. g. guereza lineages clustered with C. g. caudatus and C. g. kikuyuensis from Kenya and northern Tanzania. These two guereza lineages diverged around 0.7 million years ago. In addition, mitochondrial sequence information does not support unequivocally a distinction of C. g. caudatus and C. g. kikuyuensis. Our findings indicate a previous biogeographic connection between the ranges of C. g. occidentalis and C. g. gallarum and a possible secondary invasion of Ethiopia by members of the C. g. guereza–C. g. caudatus–C. g. kikuyuensis clade. Given these phylogenetic relationships, our study supports the two-taxa hypothesis, making C. g. gallarum an Ethiopian endemic, and, in combination with the taxon's very restricted range, makes it one of the most endangered subspecies of black-and-white colobus.
- Article
(2211 KB) - Full-text XML
-
Supplement
(494 KB) - BibTeX
- EndNote
The genus Colobus comprises five species: C. satanas, C. vellerosus, C. polykomos, C. angolensis, and C. guereza. The internal taxonomy of the genus is based on morphological features, mainly the extent of the white body markings and the proportion of white fur on the tail and the size of the tail brush (Lydekker, 1905; Schwarz, 1929; Rahm, 1970; Hull, 1978), but also on other traits, such as the acoustic structure of loud calls (“roars”) of adult males (Oates and Trocco, 1983; Oates et al., 2000). A study based on mitochondrial DNA (mtDNA) largely supports the morphology-based phylogeny with one exception: morphologically C. guereza appears to be the sister taxon of C. vellerosus (Oates and Trocco, 1983), while the molecular study suggests a closer relationship between C. polykomos and C. vellerosus (Ting, 2008), which makes more sense biogeographically because these two taxa occur parapatrically in western Africa with a possible broad hybrid zone between the Sassandra and Bandama rivers (Groves et al., 1993; Gonedelé Bi et al., 2006). Ting (2008) estimated divergence ages among species between 3.5 and 0.2 million years ago (Ma), with C. satanas diverging first followed by C. angolensis, C. guereza, and most recently C. vellerosus and C. polykomos.
C. guereza is the most widely distributed species of the genus with a more or less continuous range from Gabon and Cameroon in the west to Uganda in the east and a more fragmented distribution in Ethiopia, Kenya, and northern Tanzania (Groves, 2007; Kingdon et al., 2008; Fashing and Oates, 2013). In eastern Africa C. guereza is often confined to higher altitudes of isolated mountain ranges, such as Mt Kilimajaro, Mt Kenya, the Aberdare Range, Mt Elgon, or the Matthews Range (Fashing and Oates, 2013).
Several subspecies of C. guereza have been described, but they are less strikingly different than those of C. angolensis (Groves, 2007). C. g. caudatus, the most south-eastern form (Fig. 3e), is very different from the north-western C. g. occidentalis, but there is a string of geographically and morphologically intermediate forms (Groves, 2007). Moreover, adaptations to high mountain environments of certain populations contribute additional morphological variation, making subspecies delineation more complicate (Carpateno and Gippoliti, 1994).
Rahm (1970) recognized nine subspecies in C. guereza, whereas Dandelot (1971) recognized only six. Hull (1979) conducted a craniometric analysis of 607 skulls (340 males; 267 females) of adult guerezas from 18 populations and recognized eight subspecies (C. g. guereza, C. g. gallarum, C. g. occidentalis, C. g. dodingae, C. g. matschiei, C. g. kikuyuensis, C. g. percivali, and C. g. caudatus; Table 1). This taxonomy was subsequently adopted by IUCN (Kingdon et al., 2008) and several other authors (Groves, 2001, 2007; Grubb et al., 2003; Anandam et al., 2013; Fashing and Oates, 2013) (see Table 1). Recently, Butynski and de Jong (2018) suggested that C. g. caudatus should be elevated to species rank because of its extreme geographic isolation and phenotypical distinctiveness. At least concerning morphology, this suggestion is not in agreement with Hull (1979), who found that C. g. gallarum is the most distinctive guereza subspecies. Based on his morphological analysis, Hull (1979) also provided a phylogeny of the subspecies of C. guereza, with C. g. gallarum as the basal lineage and C. g. occidentalis as the most recent. However, to date, no molecular study has been carried out to further clarify the internal phylogeny of C. guereza.
Table 1History of the taxonomic classification of Colobus guereza.
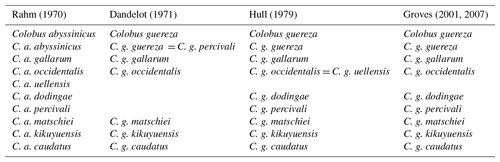

Grubb et al. (2003), Fashing and Oates (2013), and Anandam et al. (2013) adopted the taxonomy proposed by Hull (1979) and Groves (2001, 2007).
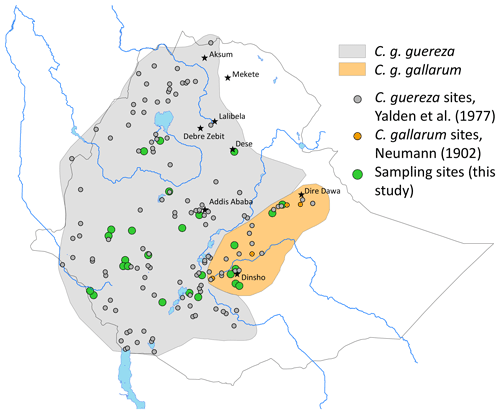
Two of the subspecies – C. g. guereza and C. g. gallarum – are possibly endemic to Ethiopia (Fashing and Oates, 2013). C. g. guereza is widely distributed from the Omo River valley to the Ethiopian Highlands west, north-west, and south-west of the Rift Valley (Yalden et al., 1977). Its southern distribution limits are not well defined, and they might extend into Kenya (Dandelot, 1971). The distribution of C. g. gallarum is even less precisely known. They are reported to be confined to the Ethiopian Highlands, east of the Rift (Gippoliti and Butynski, 2008), but Dandelot (1971) wrote that they occur north-east of the Rift Valley (which is probably wrong) and in the north-eastern part of Oromia, where they occur south-east of the Rift Valley. Fashing and Oates (2013) wrote that C. g. gallarum occurs in the Ethiopian Highlands east of the Rift Valley, whereas Carpaneto and Gippoliti (1994) noted striking differences among guerezas east of the Rift Valley, i.e. among guerezas from the Harenna Forest of the Bale Mts and guerezas from the type locality of C. g. gallarum.
Earliest records, including a description of the holotype (Colobus gallarum Neumann, 1902), were provided by Neumann (1902, p. 3: ”Berge im Quellgebiet des Webbi Shebeli; Gara Mulata bei Harar und Djaffa-Berge im Arussi-Land, ferner Wälder bei Burka auf der Straße von Harar nach Adis Abeba”; ”from mountains near the headwaters of the Webi Shabeelle (Arussi Mts?), from Gara Mulata near Harar, the Djaffa Mountains in Arussi country, and from the forests near Burka along the road from Harar to Addis Abeba”, transl. D. Zinner).
The guerezas of Ethiopia occupy a wide variety of forest habitats such as tropical deciduous forest, montane Juniperus and Hagenia forest, and riverine forests at altitudes between 400 and 3300 m (Yalden et al., 1977). Since they are not confined to high-altitude habitats, the low-lying Rift Valley most likely does not constitute a dispersal barrier as for other Ethiopian highland species, e.g. Ethiopian wolves Canis simensis (Gottelli et al., 2004), geladas Theropithecus gelada (Shotake et al., 2016; Zinner et al., 2018a), and several rodent and frog species (Bryjaa et al., 2018; Evans et al., 2011; Freilich et al., 2016).
The classification of C. g. guereza and C. g. gallarum was mainly based on craniometric differences (Hull, 1979) and on colour differences in their tail bases (Rahm, 1970; Fashing and Oates, 2013). In contrast, Yalden et al. (1977) considered at least the tail colour as a clinal trait, and he noted that no geographic barrier exists between the two populations; thus, he did not accept the subspecific classification. As for the other subspecies of C. guereza, no molecular analysis is available for the two Ethiopian taxa.
The aim of our study was a phylogenetic reconstruction of mitochondrial lineages of Ethiopian C. guereza to test whether the mtDNA-phylogeny is congruent with the taxonomic distinction between C. g. guereza and C. g. gallarum as suggested by, e.g., Hull (1979) or whether it supports the clinal hypothesis of Yalden et al. (1977).
2.1 Ethical statement
Sample collection complied with the laws of Ethiopia and Germany and with the guidelines of the International Primatological Society. During sampling of faecal material, no animals were harmed or disturbed.
2.2 Sample collection
We non-invasively collected fresh faecal samples at 26 sites in the range of C. guereza (Fig. 1 and Table S1 in the Supplement). Geographic coordinates of each sample were assigned at the time of collection by GPS. Faecal samples were collected and stored following the two-step protocol of Nsubuga et al. (2004) and Roeder et al. (2004). Samples were stored at ambient temperature for up to 3 months in the field and at −20 ∘C upon arrival in the laboratory of the German Primate Center (DPZ).
2.3 Laboratory methods
We extracted total genomic DNA using the First DNA all tissue kit (Gen-Ial) according to the manufacturer's protocols, with minor modifications as outlined in Kalbitzer et al. (2016). After extraction, DNA concentration was measured with a NanoDrop ND-1000 spectrophotometer (Peqlab) and extracts were stored at −20 ∘C until further processing.
We amplified and sequenced two mitochondrial fragments, the complete cytochrome b gene (cytb; 1140 bp) and a region spanning a portion of the NADH dehydrogenase subunit 3 gene, the tRNA for arginine, the complete NADH dehydrogenase subunit 4L gene, and a portion of the NADH dehydrogenase subunit 4 gene (NADH; 790 bp). We focused on these two markers because for both, several orthologous sequences from Colobus are available in GenBank for comparisons.
Cytb was amplified via two overlapping PCR products with sizes of 727 and 663 bp, while NADH was amplified via a single PCR product with a size of 873 bp (for primers see Table S2). To minimize the risk of amplifying nuclear mitochondrial-like sequences (numts), primers were specifically designed for C. guereza on the basis of available sequence data in Genbank. We conducted PCR reactions in a total volume of 30 µL containing a final concentration of 0.33 µmol of each primer, 3 mmol MgCl2, 0166 mmol dNTPs, 1× buffer, 1 U Biotherm Taq DNA polymerase (Genecraft) and 100 ng total genomic DNA. Cycling conditions consisted of pre-denaturation for 2 min at 96 ∘C, followed by 40 cycles, each with denaturation for 1 min at 96 ∘C, annealing for 1 min at 50 ∘C for both cytb fragments and 60 ∘C for NADH, and extension for 1 min at 72 ∘C. At the end, a final extension step for 5 min at 72 ∘C was added. To check for PCR performance, aliquots of the PCR products were run on 1 % agarose gels. PCR products were cleaned with ExoSAP-IT™ Express PCR Product Cleanup Reagent from ThermoFisher and subjected to sequencing using the amplification primers. Sanger sequencing was conducted at GATC Biotech. Sequence electropherograms were checked by eye with 4Peaks 1.8 (https://www.nucleobytes.com, last access: 18 December 2018) and sequences were assembled and manually edited in SeaView 4.4.0 (Gouy et al., 2010). SeaView was also used to check for the correct translation of protein-coding sequences into amino acid sequences.
To reduce cross-sample contamination, all working steps (DNA extraction, PCR set-up, PCR amplification, gel electrophoresis, and PCR product purification) were conducted in separate laboratories. Work benches were cleaned with 10 % bleach and new gloves were used for each sample. Further, PCR controls (without template DNA) were routinely conducted and procedures were repeated for 10 % of randomly selected samples.
2.4 Data analysis
We expanded our dataset with orthologous Colobus sequences from GenBank (Table S1). We generated three datasets: NADH (103 sequences), cytb (79 sequences), and the concatenated NADH and cytb (59 sequences). The number of sequences for each marker differed because (1) for the out-group and most of the non-Ethiopian C. guereza taxa, orthologues sequences were not available in GenBank and (2) we were not able to sequence NADH and cytb from every sample.
For phylogenetic tree reconstructions, identical haplotypes were removed, resulting in datasets with 53 (NADH), 34 (cytb), and 33 (concatenated) sequences. Phylogenetic trees were constructed with maximum-likelihood (ML) methods using IQ-Tree 1.5.2 (Nguyen et al., 2015) and Bayesian inferences with MrBayes 3.2.6 (Ronquist et al., 2012). The best-fit models for each dataset were determined with ModelFinder (Chernomor et al., 2016; Kalyaanamoorthy et al., 2017) in IQ-Tree under the Bayesian information criterion (BIC). The ML tree was reconstructed with 10 000 ultra-fast bootstrap (BS) replicates (Minh et al., 2013), while the Bayesian tree was obtained from a Markov chain Monte Carlo (MCMC) run with 10 million generations, sampling every 1000 generations. For the Bayesian reconstruction, we checked convergence of all parameters and the adequacy of a 25 % burn-in by assessing the uncorrected potential scale reduction factor (PSRF) (Gelman and Rubin, 1992). Posterior probabilities (PPs) for nodes and a phylogram with mean branch lengths were calculated from the posterior density of trees using MrBayes. Phylogenetic trees were visualized in FigTree 1.4.2 (http://tree.bio.ed.ac.uk/software/figtree/, last access: 18 December 2018).
Divergence times were estimated with a Bayesian approach as implemented in BEAST 2.5.0 (Bouckaert et al., 2014). We performed two independent analyses for each dataset, each with 25 million generations and tree and parameter sampling occurring every 1000 generations. For the analyses, we assumed a relaxed log-normal clock model and applied a coalescent constant population prior for branching rates. As no reliable information from the fossil record is available to calibrate the molecular clock, we used estimated divergence times based on molecular data (Ting, 2008). For both datasets, we set age constraints using a normal prior. For the NADH dataset, we constrained the split between C. satanas and the other Colobus species at a mean of 3.5 with sigma 0.36, translating into a median of 3.5 Ma and a 95 % highest probability density (HPD) of 2.8–4.2 Ma. For the cytb and concatenated datasets, we constrained the split between C. angolensis and C. guereza at a mean of 2.1 with sigma 0.26, translating into a median of 2.1 Ma and a 95 % HPD of 1.6–2.6 Ma. The adequacy of a 10 % burn-in and convergence of all parameters was assessed by inspecting the trace of the parameters across generations using Tracer 1.6 (http://beast.bio.ed.ac.uk/Tracer, last access: 18 December 2018). Sampling distributions of the run were combined with LogCombiner 2.5.0, and the tree with mean node heights was summarized with TreeAnnotator 2.5.0 using a burn-in of 10 %. The trees were visualized in FigTree.
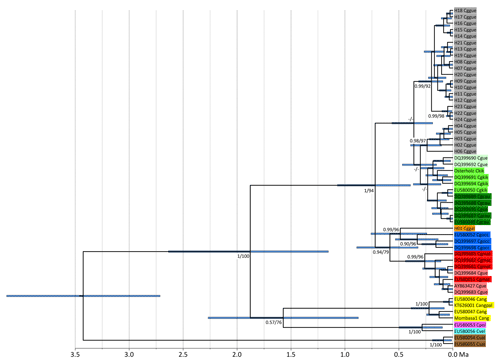
Figure 2Ultrametric tree showing phylogenetic relationships and divergence times of Colobus mtDNA lineages (based on 790 bp of NADH). Tip labels refer to Colobus haplotypes (see Table S1). Csat: Colobus satanas; Cvel: C. vellerosus; Cpol: C. polykomos; Cang: C. angolensis; Cgocc: C. guereza occidentalis; Cggal: C. g. gallarum; Cgmat: C. g. matschiei; Cggue: C. g. guereza; Cgkik: C. g. kikuyuensis; Cgcau: C. g. caudatus; Cgue: C. guereza ssp. Node labels refer to ML BS and Bayesian PP support values. The timescale below the tree indicates million years BP.
For NADH we retrieved 74 sequences containing 24 haplotypes, and for cytb we retrieved 76 sequences including 31 haplotypes, resulting in 56 concatenated sequences including 30 haplotypes. Independently of the markers used and algorithms applied, the phylogenetic reconstructions show the same tree topology (Figs. 2, S1–S3 in the Supplement). The specimens of the eastern part of the original C. g. gallarum range have highly distinct haplotypes, compared to specimens from the range of C. g. guereza. The phylogeny based on NADH (for which we were able to include sequences of several other Colobus taxa) revealed that the supposed C. g. gallarum haplotypes cluster with supposed C. g. occidentalis (Fig. 2). The combined C. g. gallarum + C. g. occidentalis clade clusters with a clade containing supposed C. g. matschiei. These three taxa form a clade which is the sister clade of a mixed clade of C. g. caudatus, C. g. kikuyuensis, and C. g. guereza. The overall Colobus phylogeny revealed the basal position of C. satanas, forming the out-group to all other Colobus linages (Fig. 2). At around 1.8 Ma a combined clade of the western African taxa (C. vellerosus and C. polykomos) and C. angolensis split off from C. guereza. Within C. guereza the first divergence appeared at around 0.7 Ma, when the lineage leading to C. g. occidentalis, C. g. gallarum, and C. g. matschiei separated from the lineage leading to C. g. caudatus, C. g. kikuyuensis and C. g. guereza. C. g. matschiei split off from the C. g. occidentalis + C. g. gallarum clade at around 0.6 Ma, which further diverged at around 0.5 Ma. C. g. caudatus and C. g. kikuyuensis separated only at around 0.2 Ma. The phylogenetic relationships among C. g. caudatus, C. g. kikuyuensis and C. g. guereza are not well resolved, most likely because they diverged relatively recently (<0.3 Ma).
The geographic pattern of the distribution of C. guereza haplotypes in Ethiopia revealed that the supposed C. g. gallarum haplotype was only found at two sites in the eastern part of the C. g. gallarum range whereas those specimens from the western part of the range (e.g. Arsi [Arusi, Arussi] Mts) cluster with C. g. guereza.
Our study reveals that mitochondrial haplotypes of C. guereza form two major clades which separated around 0.7 Ma. One clade contains haplotypes of C. g. occidentalis, C. g. gallarum, and C. g. matschiei, the other haplotypes of C. g. caudatus, C. g. kikuyuensis, and C. g. guereza. The phylogenetic relationships among the latter three taxa are not resolved, which is most likely a result of their recent divergence where lineage sorting is still incomplete. A complete phylogenetic analysis of all supposed taxa of C. guereza was not possible because haplotypes of C. g. dodingae and C. g. percivali were not available.
Two major mitochondrial lineages occur in Ethiopia, and their geographic distribution largely corresponds to the geographic ranges of C. g. guereza and C. g. gallarum (Fig. 1). However, the actual range of the C. g. gallarum haplotype is much smaller than the range of C. g. gallarum as depicted in, e.g., Fashing and Oates (2013). The C. g. gallarum haplotype was found only in two local populations in the eastern part of the assumed range of C. g. gallarum, whereas in the western part of the range (Arussi Mts), we found only C. g. guereza haplotypes. This supports the notion of Carpaneto and Gippoliti (1994) that guerezas of the western part (e.g. Harenna Forest) are phenotypically different from C. g. gallarum. The Rift Valley does not constitute a boundary between C. g. gallarum and C. g. guereza haplotypes, contradicting the notion by Fashing and Oates (2013) that C. g. guereza is confined to the west of the Rift Valley.
4.1 Taxonomy
In primates, as in other taxonomic groups, taxon delimitation based on mitochondrial sequence information often remains unreliable if the respective divergences or radiations are relatively recent. Often mitochondrial clades mirror biogeographic patterns rather than taxonomy (e.g. baboons; Zinner et al., 2009, 2015, 2018b). Since our study was based on mitochondrial markers, we thus have to be cautious when discussing possible taxonomic inferences.
However, results of our study principally support the two-taxa hypothesis for Ethiopian black-and-white colobus. Notably the two Ethiopian mitochondrial lineages' membership of to two distinct C. guereza clades that separated at around 0.7 Ma somehow indicates independent evolutionary histories of the putative subspecies. This divergence age is even more remarkable as the divergence ages among C. g. guereza, C. g. caudatus, and C. g. kikuyuensis are much younger (<0.3 Ma). Also, the phylogenetic relationships among the three taxa, in particular between C. g. caudatus and C. g. kikuyuensis are not well resolved and only weakly supported (Fig. S1a and S1b). Therefore, mitochondrial sequence information also does not provide evidence for C. g. caudatus being the most distinctive form of black-and-white colobus subspecies and thus, do not support the proposal by Butynski and de Jong (2018) to elevate C. g. caudatus to a separate species (C. caudatus). However, at the moment our taxonomic inferences regarding non-Ethiopian C. guereza have to be preliminary since the geographic provenance of several mtDNA sequences is from zoo animals, and we have to rely on the taxonomic classification provided by the authors of the respective sequences.
Since hybridization and introgression is common among various primate taxa (Zinner et al., 2011; Cortes-Ortiz et al., 2019), the Ethiopian guerezas might also be affected, which would partially explain the different views regarding their taxonomy and delineation of subspecies (Yalden et al., 1977; Fashing and Oates, 2013). A thorough study based on genomic data and a reassessment of their morphology would help to shed light on the internal phylogeny of C. guereza.
4.2 Phylogeography
The close phylogenetic relationship of the eastern Ethioipian C. g. gallarum with the central African lowland C. g. occidentalis was unexpected. C. guereza is ecologically relatively flexible (Fashing, 1999; Fashing and Oates, 2013; Harris and Chapman, 2007) but, as an arboreal species, depends on forest or woodland. Given the historical changes in climate and forest cover in Africa (deMenocal, 2004; Trauth et al., 2009; Hoag and Svenning, 2017), one can assume that ancestral C. guereza populations became isolated in forest refugia under unfavourable conditions, whereas forests and C. guereza populations became reconnected under favourable conditions.
Assuming an origin of C. guereza in the northern forest belt of central Africa, a possible phylogegeographic scenario (Fig. 3) could have started with a range expansion into the forests of north-east Africa and Ethiopia, followed by a first divide into the more central and northern ancestral population of the C. g. occidentalis + C. g. matschiei + C. g. gallarum clade and more southern population of C. g. guereza + C. g. kikuyuensis + C. g. caudatus clade. Both populations further radiated into the respective subspecies. Under unfavourable conditions, C. g. gallarum became isolated in the extreme eastern part of the range. The gap between the eastern and more western and central distribution was then probably filled by a range expansion of the southern clade. In order to find support for such a scenario, genetic analyses of the missing populations of C. g. dodingae and C. g. percivali would be helpful. One could expect clustering of C. g. dodingae with C. g. occidentalis or C. g. matschiei, which would narrow the gap towards C. g. gallarum and clustering of C. g. percivali with C. g. kikuyuensis or C. g. guereza, supporting a south–north expansion into the gap.
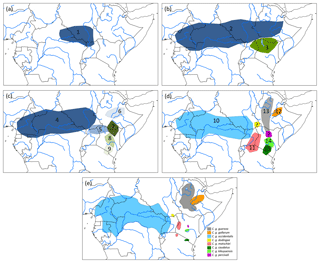
Figure 3Phylogeographic scenario for Colobus guereza. (a) Ancestral population (1) in the northern forest belt; (b) distribution of two subpopulations after the first split of the ancestral lineage into ancestral populations of C. g. occidentalis + C. g. matschiei + C. g. gallarum (2) and C. g. caudatus + C. g. kikuyuensis + C. g. guereza (3); (c) further split of population (2) into ancestral C. g. occidentalis (4), C. g. matschiei (5), and C. g. gallarum (6) populations and split of population (3) into ancestral C. g. guereza (7), C. g. kikuyuensis (8), and C. g. caudatus (9) populations; (d) further differentiation of ancestral populations and northern range expansion of C. g. guereza into Ethiopia; the phylogenetic relationships of C. g. percivali and C. g. dodingae have to currently remain open because relevant mtDNA sequences are not available; (e) current distribution of C. guereza taxa modified after Fashing and Oates (2013).
4.3 Conservation
If the distinctiveness of the populations carrying C. g. gallarum haplotypes is supported by morphological and nuclear characters, C. g. gallarum is most likely the most threatened subspecies of C. guereza. At the moment the C. g. gallarum haplotypes are only found at two sites where small populations may subsist. Since the forests in the supposed former range of C. g. gallarum have been heavily reduced, the outlook for this taxon is not promising (Gippoliti and Butynski, 2008). In contrast, C. g. guereza is still widely distributed in Ethiopia. Since both Ethiopian guereza taxa are closely related, there is a certain danger of hybridization if they meet. It would probably be interesting, whether genetic traces of earlier hybridization can be found in C. g. gallarum. At the moment, the C. g. gallarum populations seem to be geographically isolated from the closest C. g. guereza population, but care should be taken to prevent the restocking of the small C. g. gallarum population with C. g. guereza individuals, even for conservation reasons (Gippoliti et al., 2018).
It also becomes obvious that, if genetic diversity and local adaptation across the complete range are to be preserved (Harcourt, 2006; Harcourt et al., 2005), several subspecies of C. guereza need particular conservation measures, including C. g. gallarum.
Our mitochondrial phylogeny revealed that two mitochondrial clades exist in Ethiopian guerezas and that their respective distributions are largely congruent with the geographic ranges of C. g. gallarum and C. g. guereza. The phylogenetic relationships of the two mitochondrial clades indicate that the two subspecies are not sister taxa. In contrast, our findings indicate a previous biogeographic connection between the ranges of C. g. occidentalis and C. g. gallarum and a possible secondary invasion of Ethiopia by members of the C. g. guereza–C. g. caudatus–C. g. kikuyuensis clade. Given these phylogenetic relationships, our study supports the two-taxa hypothesis, and in combination with its very restricted range, makes C. g. gallarum one of the most endangered subspecies of black-and-white colobus. However, to fully understand the phylogenetic relationship among C. guereza populations, a reassessment of their morphology might be helpful, and in any case, a study using nuclear markers is needed.
Sequence data are available from NCBI GenBank (https://www.ncbi.nlm.nih.gov/).
The supplement related to this article is available online at: https://doi.org/10.5194/pb-6-7-2019-supplement.
DZ, AA, DT, and CR conceived the study. SD, DT, and AM collected the samples in Ethiopia. NCS and AB supported sample collection. AA, DT, DZ, and CR produced and analysed the data, and DZ, CR and AA wrote the paper.
The authors declare that they have no conflict of interest.
We thank Christiane Schwarz for her help in the lab and the Ethiopian Wildlife Conservation Authority for providing permits to conduct the field work and to collect the samples. We also thank two reviewers (Sarah Elton and Spartaco Gippoliti) for their valuable comments. We are also grateful to Rufford Small Grants for Nature Conservation, Nature and Biodiversity Conservation Union (NABU) and the Alexander von Humboldt Foundation for supporting our project.
This paper was edited by Sarah Elton and reviewed by Spartaco Gippoliti and Sarah Elton.
Anandam, M. V., Bennett, E. L., Davenport, T. R. B., Davies, N. J., Detwiler, K. M., Engelhardt, A., Eudey, A. A., Gadsby, E. L., Groves, C. P., Healy, A., Karanth, K. P., Molur, S., Nadler, T., Richardson, M. C., Riley, E. P., Roos, C., Rylands, A. B., Sheeran, L. K., Ting, N., Wallis, J., Waters, S. S., Whittaker, D. J., and Zinner, D.: Species accounts of Cercopithecidae, in: Handbook of the Mammals of the World Vol 3 Primates, edited by: Mittermeier, R. A., Rylands, A. B., and Wilson, D. E., Lynx Edicions, Barcelona, 628–753, 2013.
Bouckaert, R., Heled, J., Kuehnert, D., Vaughan, T., Wu, C. H., Xie, D., Suchard, M. A., Rambaut, A., and Drummond, A. J.: BEAST 2: a software platform for Bayesian evolutionary analysis, PLoS Comput. Biol., 10, e1003537, https://doi.org/10.1371/journal.pcbi.1003537, 2014.
Bryjaa, J., Kostin, D., Meheretu, Y., Šumbera, R., Bryjová, A., Kasso, M., Mikula, O., and Lavrenchenko, L. A.: Reticulate Pleistocene evolution of Ethiopian rodent genus along remarkable altitudinal gradient, Mol. Phylogenet. Evol., 118, 75–87, 2018.
Butynski, T. M. and de Jong, Y. A.: Geographic range, taxonomy, and conservation of the Mount Kilimanjaro guereza colobus monkey (Primates: Cercopithecidae: Colobus), Hystrix, 29, 81–85, 2018.
Carpaneto, G. M. and Gippoliti, S.: Primates of the Harenna Forest, Ethiopia, Primate Conserv., 11, 12–15, 1994.
Chernomor, O., von Haeseler, A., and Minh, B. Q.: Terrace aware data structure for phylogenomic inference from supermatrices, Syst. Biol., 65, 997–1008, 2016.
Cortes-Ortiz, L., Roos, C., and Zinner, D.: Introduction to Special Issue on Primate Hybridization and Hybrid Zones, Int. J. Primatol., 40, 1–8, 2019.
Dandelot, P.: Order Primates, in: The Mammals of Africa. An Identification Manual, Part 3, edited by: Meester, J. and Setzer H. W., Smithsonian Institution Press, Washington, D.C., 1–43, 1971.
deMenocal, P. B.: African climate change and faunal evolution during the Pliocene-Pleistocene, Earth Planet. Sc. Lett., 220, 3–24, 2004.
Evans, B. J., Bliss, S. M., Mendel, S. A., and Tinsley, R. C.: The Rift Valley is a major barrier to dispersal of African clawed frogs (Xenopus) in Ethiopia, Mol. Ecol., 20, 4216–4230, 2011.
Fashing, P. J.: The Behavioral Ecology of an African Colobine Monkey: Diet, Range Use, and Patterns of Intergroup Aggression in Eastern Black and White Colobus Monkeys (Colobus guereza), PhD thesis, Columbia University, New York, 1999.
Fashing, P. J. and Oates, J. F.: Colobus guereza guereza colobus, in: Mammals of Africa Volume II: Primates, edited by: Butynski, T. M., Kingdon, J., and Kalina, J., Bloomsbury, London, 111–119, 2013.
Freilich, X., Anadón, J. D., Bukala, J., Calderon, O., Chakraborty, R., and Boissinot, S.: Comparative phylogeography of Ethiopian anurans: impact of the Great Rift Valley and Pleistocene climate change, BMC Evol. Biol., 16, 206, https://doi.org/10.1186/s12862-016-0774-1, 2016.
Gelman, A. and Rubin, D. B.: Inference from iterative simulation using multiple sequences (with discussion), Stat. Sci., 7, 457–511, 1992.
Gippoliti, S. and Butynski, T. M.: Colobus guereza ssp. gallarum, The IUCN Red List of Threatened Species, e.T5150A11118023, https://doi.org/10.2305/IUCN.UK.2008.RLTS.T5150A1111802 3.en (last access: 2 January 2019), 2008.
Gippoliti, S., Cotterill, F. P. D., Groves, C. P., and Zinner, D.: Poor taxonomy and genetic rescue are possible co-agents of silent extinction and biogeographic homogenization among ungulate mammals, Biogeographia, 33, 41–54, 2018.
Gonedelé Bi, S. E., Zinner, D., Koné, I., Goné Bi, Z., Akpatou, B, Koffi Bené, J. C., Sangaré, A., and Boesch, C.: A West African black-and-white colobus monkey, Colobus polykomos dollmani Schwarz, 1927, facing extinction, Primate Conserv., 21, 55–61, 2006.
Gottelli, D., Marino, J., Sillero-Zubiri, C., and Funk, S. M.: The effect of the last glacial age on speciation and population genetic structure of the endangered Ethiopian wolf (Canis simensis), Mol. Ecol., 13, 2275–2286, 2004.
Gouy, M., Guindon, S., and Gascuel, O.: SeaView Version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building, Mol. Biol. Evol., 27, 221–224, 2010.
Groves, C. P.: Primate Taxonomy, Smithsonian Institution Press, Washington, D.C., 2001.
Groves, C. P.: The taxonomic diversity of the Colobinae of Africa, J. Anthropol. Sci., 85, 7–34, 2007.
Groves, C. P., Angst, R., and Westwood, C.: The status of Colobus polykomos dollmani Schwarz, Int. J. Primatol., 14, 573–586, 1993.
Grubb, P., Butynski, T. M., Oates, J. F., Bearder, S. K., Disotell, T. R., Groves, C. P., and Struhsaker, T. T.: Assessment of the diversity of African primates, Int. J. Primatol., 24, 1301–1313, 2003.
Harcourt, A. H.: Rarity in the tropics: biogeography and macroecology of the primates, J. Biogeogr., 33, 2077–2087, 2006.
Harcourt, A. H., Coppeto, S. A., and Parks, S. A.: The distribution-abundance (density) relationship: its form and causes in a tropical mammal order, Primates, J. Biogeogr., 32, 565–579, 2005.
Harris, T. R. and Chapman, C. A.: Variation in diet and ranging of black and white colobus monkeys in Kibale National Park, Uganda, Primates, 48, 208–221, 2007.
Hoag, C. and Svenning, J. C.: African environmental change from the Pleistocene to the Anthropocene, Annu. Rev. Environ. Resour., 42, 27–54, 2017.
Hull, D. B.: Aberrations in the coat colour patterns of black and white colobus monkeys, Afr. J. Ecol., 16, 21–27, 1978.
Hull, D. B.: A craniometric study of the black and white Colobus Illiger 1811 (Primates: Cercopithecoidea), Am. J. Phys. Anthropol., 51, 163–182, 1979.
Kalbitzer, U., Roos, C., Kopp, G. H., Butynski, T. M., Knauf, S., Zinner, D., and Fischer, J.: Insights into the genetic foundation of aggression in Papio and the evolution of two length-polymorphisms in the promoter regions of serotonin-related genes (5-HTTLPR and MAOALPR) in Papionini, BMC Evol. Biol., 16, 121, https://doi.org/10.1186/s12862-016-0693-1, 2016.
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A., and Jermiin, L. S.: ModelFinder.: fast model selection for accurate phylogenetic estimates, Nat. Methods, 14, 587–589, 2017.
Kingdon, J., Struhsaker, T., Oates, J. F., Hart, J., and Groves, C. P.: Colobus guereza, The IUCN Red List of Threatened Species, e. T5143A11116447, https://doi.org/10.2305/IUCN.UK.2008.RLTS.T5143A1111644 7.en (last access: 2 January 2019), 2008.
Lydekker, R.: Colour evolution in guereza monkeys, Proc. Zool. Soc. Lond., 1905, 325–329, 1905.
Minh, B. Q., Nguyen, M. A. T., and von Haeseler, A.: Ultrafast approximation for phylogenetic bootstrap, Mol. Biol. Evol., 30, 1188–1195, 2013.
Neumann, O.: Neue nordost- und ostafrikanische Säugethiere, Sitzungs-Bericht der Gesellschaft naturforschender Freunde zu Berlin, 3, 3–59, 1902.
Nguyen, L. T., Schmidt, H. A., von Haeseler, A., and Minh, B. Q.: IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies, Mol. Biol. Evol., 32, 268–274, 2015.
Nsubuga, A. M., Robbins, M. M., Roeder, A. D., Morin, P. A., Boesch, C., and Vigilant, L.: Factors affecting the amount of genomic DNA extracted from ape faeces and the identification of an improved sample storage method, Mol. Ecol., 13, 2089–2094, https://doi.org/10.1111/j.1365-294X.2004.02207.x, 2004.
Oates, J. F. and Trocco, T. F.: Taxonomy and phylogeny of black-and-white colobus monkeys: inferences from an analysis of loud call variation, Folia Primatol., 40, 83–113, 1983.
Oates, J. F., Bocian, C. M., and Terranova, C. J.: The loud calls of black-and-white colobus monkeys: their adaptive and taxonomic significance in light of new data, in: Old World Monkeys, edited by: Whitehead, P. F. and Jolly, C. J., Cambridge University Press, Cambridge, 431–452, 2000.
Rahm, U.: Ecology, zoogeography, and systematics of some African forest monkeys, in: Old World Monkeys, edited by: Napier, J. R. and Napier, P. H., Academic Press, New York, 589–626, 1970.
Roeder, A. D., Archer, F. I., Poiner, H. N., and Morin, P. A.: A novel method for collection and preservation of faeces for genetic studies, Mol. Ecol. Notes, 4, 761–764, https://doi.org/10.1111/j.1471-8286.2004.00737.x, 2004.
Ronquist, F., Teslenko, M., van der Mark, P., Ayres, D. L., Darling, A., Höhna, S., Larget, B., Liu, L., Suchard, M. A., and Huelsenbeck, J. P.: MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space, Syst. Biol., 61, 539–542, 2012.
Rüppell, E.: Neue Wirbelthiere zu der Fauna von Abbyssinien gehörig, Säugethiere, Siegmund Schmerber, Frankfurt-am-Main, 1–4, 1835.
Schwarz, E.: On the local races and distribution of the black and white colobus monkeys, Proc. Zool. Soc. Lond., 3, 585–598, 1929.
Shotake, T., Saijuntha, W., Agatsuma, T., and Kawamoto, Y.: Genetic diversity within and among gelada (Theropithecus gelada) populations based on mitochondrial DNA analysis, Anthropol. Sci., 124, 157–167, 2016.
Ting, N.: Mitochondrial relationships and divergence dates of the African colobines: evidence of Miocene origins for the living colobus monkeys, J. Hum. Evol., 55, 312–325, 2008.
Trauth, M. H., Larrasoaña, J. C., and Mudelsee, M.: Trends, rhythms and events in Plio-Pleistocene African climate, Quat. Sci. Rev., 28, 399–411, 2009.
Yalden, D. W., Largen, M. J., and Kock, D.: Catalogue of the mammals of Ethiopia. Part 3: Primates, Ital. J. Zool. NS, suppl. 9, 1–52, 1977.
Zinner, D., Groeneveld, L. F., Keller, C., and Roos, C.: Mitochondrial phylogeography of baboons (Papio spp.) – Indication for introgressive hybridization?, BMC Evol. Biol., 9, 83, https://doi.org/10.1186/1471-2148-9-83, 2009.
Zinner, D., Arnold, M. L., and Roos, C.: The strange blood: Natural hybridization in primates, Evol. Anthropol., 20, 96–103, 2011.
Zinner, D., Keller, C., Nyahongo, J. W., Butynski, T. M., de Jong, Y. A., Pozzi, L., Knauf, S., Liedigk, R., and Roos, C.: Distribution of mitochondrial clades and morphotypes of baboons Papio spp. (Primates: Cercopithecidae) in Eastern Africa, J. East Afr. Nat. Hist., 104, 143–168, 2015.
Zinner, D., Atickem, A., Beehner, J. C., Bekele, A., Bergman, T. J., Burke, R., Dolotovskaya, S., Fashing, P. J., Gippoliti, S., Knauf, S., Knauf, Y., Mekonnen, A., Moges, A., Nguyen, N., Stenseth, N. C., and Roos, C.: Phylogeography and mitochondrial DNA diversity in geladas (Theropithecus gelada), PLoS ONE, 13, e0202303, https://doi.org/10.1371/journal.pone.0202303, 2018a.
Zinner, D., Chuma, I. S., Knauf, S., and Roos, C.: Inverted intergeneric introgression between critically endangered kipunjis (Rungwecebus kipunji) and yellow baboons (Papio cynocephalus) in two disjunct populations, Biol. Lett., 14, 20170729, https://doi.org/10.1098/rsbl.2017.0729, 2018b.